Choose The Right Viral Vector: Lentivirus, Adenovirus, AAV or Retrovirus?
The selection of an appropriate viral vector constitutes a critical decision in gene delivery workflows, directly influencing transduction efficiency, transgene expression dynamics, biosafety requirements, and therapeutic applicability. While lentivirus, adenovirus, adeno-associated virus (AAV), and retrovirus represent the four predominant platforms in research and clinical settings, their distinct virological architectures, host cell interactions, and manufacturing constraints render each suitable for specific experimental contexts. This review provides a systematic, mechanism-based comparison to inform vector selection strategy.
| Lentivirus | Adenovirus | AAV | Retrovirus (γ-retrovirus) | |
|---|---|---|---|---|
| Genome | (+) ssRNA (diploid, ~9.2 kb) | dsDNA (~36 kb) | ssDNA (~4.7 kb) | (+) ssRNA (~8.3 kb) |
| Envelope | Yes (lipid bilayer with glycoproteins) | No (naked icosahedral capsid) | No (naked icosahedral capsid) | Yes (lipid bilayer) |
| Capsid symmetry | Conical (HIV-1-derived) | Icosahedral (T=25) | Icosahedral (T=1) | Icosahedral |
| Wild-type pathogenicity | HIV-1 (attenuated in vector) | Respiratory/gastrointestinal infections | Non-pathogenic (dependent on helper virus) | Oncogenic (e.g., MLV) |
| Biosafety level | BSL-2+ | BSL-2 | BSL-1 | BSL-2 |
Molecular Mechanisms of Gene Delivery
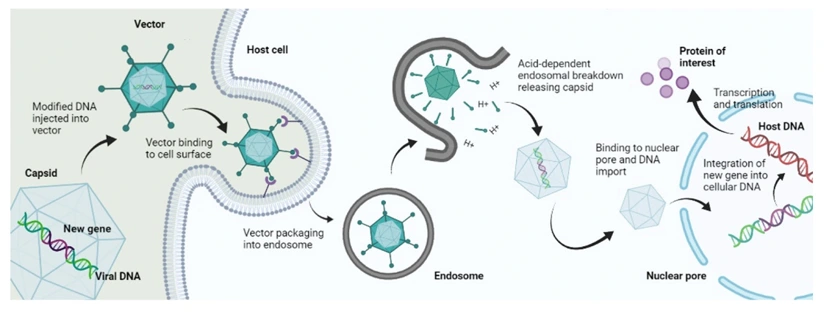
Lentivirus: Integrative, Broad-Tropism Delivery
Lentiviral vectors (primarily HIV-1-derived) enter target cells via receptor-mediated endocytosis following VSV-G pseudotyping or alternative envelope glycoprotein binding. Following cytoplasmic entry:
- Reverse transcription: Viral RNA is reverse-transcribed into double-stranded DNA by virion-packaged reverse transcriptase.
- Nuclear import: The pre-integration complex (PIC), comprising viral cDNA, integrase (IN), matrix (MA), and Vpr proteins, actively traverses the intact nuclear envelope of non-dividing cells-a defining feature distinguishing lentiviruses from conventional retroviruses.
- Chromosomal integration: Integrase catalyzes irreversible insertion of viral cDNA into host chromatin, preferentially at transcriptionally active loci.
Key implication: Stable, long-term transgene expression; however, insertional mutagenesis risk necessitates safety monitoring in clinical applications.
Adenovirus: Episomal, High-Capacity Transduction
Adenoviral vectors bind coxsackie-adenovirus receptor (CAR) and integrins (αvβ3/αvβ5), triggering clathrin-mediated endocytosis:
- Endosomal escape: Capsid protein VI mediates lysis of endosomal membrane.
- Nuclear trafficking: DNA-containing capsid docks at nuclear pore complexes; viral DNA enters the nucleoplasm.
- Episomal persistence: Viral DNA remains extrachromosomal, with no integration into host genome.
Key implication: Robust, transient transgene expression (days to weeks); minimal genotoxicity; potent inflammatory response due to viral protein expression and innate immune sensing.
AAV: Episomal, Low-Immunogenic Precision
AAV requires co-receptor binding (e.g., AAVR) and primary receptor engagement (glycans, integrins) for cellular entry:
- Endocytosis and trafficking: Clathrin-mediated or macropinocytic entry; cytoplasmic trafficking via endosomal compartments.
- Nuclear entry: Uncoating at nuclear periphery; ssDNA conversion to dsDNA by host DNA polymerase or annealing of complementary strands from co-infecting particles.
- Episomal persistence: Predominantly forms concatemeric episomes; rare, site-specific integration at AAVS1 locus (chromosome 19) in the presence of Rep protein.
Key implication: Long-term expression (months to years in non-dividing tissues); minimal immunogenicity; limited packaging capacity (~4.7 kb).
Retrovirus (γ-retrovirus): Integrative, Division-Dependent Delivery
Murine leukemia virus (MLV)-based vectors share the lentiviral life cycle but lack nuclear import machinery:
- Cell entry: Receptor-mediated (e.g., ecotropic, amphotropic, or pantropic envelope variants).
- Reverse transcription and PIC formation: Analogous to lentivirus.
- Nuclear access restricted to mitosis: Nuclear envelope disassembly during M phase is required for PIC chromosomal access.
- Integration: Integrase-mediated insertion; preference for transcription start sites and CpG islands.
Key implication: Stable integration but exclusively in dividing cells; significant insertional oncogenesis risk (demonstrated in X-SCID gene therapy trials).
Comparative Functional Characteristics
| Feature | Lentivirus | Adenovirus | AAV | Retrovirus |
|---|---|---|---|---|
| Packaging capacity | ~8 kb | ~36 kb (gutless: ~36 kb) | ~4.7 kb | ~8 kb |
| Integration | Yes (stable, random) | No (episomal) | Rarely (predominantly episomal) | Yes (stable, random) |
| Expression duration | Long-term (months to years) | Transient (days to weeks) | Long-term (months to years in non-dividing cells) | Long-term (months to years) |
| Dividing cells | Yes | Yes | Yes (but preferentially non-dividing for persistence) | Required |
| Non-dividing cells | Yes | Yes | Yes | No |
| Transduction efficiency | High | Very high | Moderate to high (serotype-dependent) | Moderate |
| Immunogenicity | Low to moderate | High (innate and adaptive) | Very low | Low to moderate |
| Inflammatory response | Minimal | Potent | Minimal | Minimal |
| Pre-existing immunity | Rare | Common (Ad5 serotype) | Common (AAV2, AAV8 in humans) | Rare |
| Clinical status | Approved (CAR-T, ex vivo gene therapy) | Approved (vaccines, oncolytic); limited gene therapy | Approved (Luxturna, Zolgensma, Hemgenix) | Limited (X-SCID historical; largely superseded) |
Tissue Tropism and Targeting Considerations
Lentivirus
- Default tropism: Broad (VSV-G pseudotyping enables entry into virtually all mammalian cell types).
- Targeting strategies: Envelope pseudotyping (e.g., RD114 for hematopoietic cells, BaEV-Rless for T cells); surface engineering (scFv display, DARPins).
Adenovirus
- Default tropism: Hepatotropic (Ad5); respiratory epithelium (Ad2/Ad5).
- Targeting strategies: Fiber knob modification (CAR-independent entry); hexon hypervariable region engineering; transcriptional targeting (tissue-specific promoters).
AAV
- Serotype-dependent tropism: Over 100 natural serotypes and engineered variants with distinct receptor usage and tissue preferences.
Retrovirus
- Default tropism: Dependent on envelope choice (ecotropic: mouse only; amphotropic: broad mammalian; pantropic: VSV-G or RD114).
- Limitation: Inability to transduce non-dividing cells restricts applications to ex vivo manipulation of proliferating populations (e.g., hematopoietic stem cells during cytokine-stimulated expansion).
Selecting the Optimal Vector
| Experimental Objective | Recommended Vector | Rationale |
|---|---|---|
| Stable, long-term expression in dividing and non-dividing cells | Lentivirus | Chromosomal integration; broad tropism; moderate immunogenicity |
| High-level, transient expression; large transgene (>8 kb) | Adenovirus | Large packaging capacity; episomal persistence; very high transduction efficiency |
| In vivo gene therapy; long-term expression in non-dividing tissues; low immunogenicity required | AAV | Minimal inflammatory response; long-term episomal persistence; diverse serotype targeting |
| Ex vivo hematopoietic stem cell modification; T cell engineering (historical) | Retrovirus | Stable integration; clinical precedent in early gene therapy (though largely replaced by lentivirus) |
| CRISPR/Cas9 knock-in (HDR-dependent) | AAV or Adenovirus | AAV: efficient HDR template delivery; Adenovirus: large donor templates possible |
| CRISPR/Cas9 knock-out (NHEJ-mediated) | Lentivirus or AAV | Lentivirus: stable Cas9/gRNA expression; AAV: transient, high-efficiency delivery |
| Oncolytic virotherapy | Adenovirus (replication-competent) | Potent lytic cycle; immunogenic cell death induction; tumor-selective replication |
| Vaccine vector (SARS-CoV-2, influenza) | Adenovirus (replication-deficient or competent) | Strong innate immune activation; efficient dendritic cell transduction |
| Neurological disease (CNS targeting) | AAV9, AAV-PHP.eB, AAVrh10 | BBB penetration (AAV9, PHP.eB); neuronal tropism; long-term expression |
| Liver-directed gene therapy | AAV8, AAV3B, Lentivirus | AAV8: highest hepatocyte transduction efficiency; Lentivirus: integration for permanent correction |
Safety and Regulatory Profiles
| Lentivirus | Adenovirus | AAV | Retrovirus | |
|---|---|---|---|---|
| Insertional mutagenesis | Moderate risk (preference for active genes) | Negligible (episomal) | Negligible (rare, site-specific if Rep present) | High risk (preference for promoters/enhancers) |
| Genotoxicity | Potential activation of proto-oncogenes | Minimal | Minimal | Documented oncogenesis (X-SCID trials) |
| Inflammatory toxicity | Low | High (cytokine release syndrome possible) | Very low (unless high dose/empty capsids) | Low |
| Pre-existing neutralizing antibodies | Rare | Common (limits efficacy) | Common (serotype-dependent) | Rare |
| Replication-competent virus (RCV) | Rigorous testing required (3-plasmid system minimizes risk) | Testing required (E1-deletion in HEK293) | No replication competence (helper-dependent) | Testing required |
Conclusion
The selection among lentivirus, adenovirus, AAV, and retrovirus is not merely a technical preference but a strategic decision grounded in the biological requirements of the target cell population, the desired duration and level of transgene expression, the acceptable safety margin, and the constraints of manufacturing scale. Lentivirus remains the platform of choice for stable genetic modification of diverse cell types, including non-dividing populations. Adenovirus excels in applications demanding high-level transient expression or large transgene capacity, albeit with significant immunogenic trade-offs. AAV has emerged as the preeminent vector for in vivo gene therapy, offering an unparalleled combination of long-term expression, low immunogenicity, and tissue-specific targeting through serotype engineering. Retrovirus, while historically significant, has been largely supplanted by lentivirus in modern applications due to its restriction to dividing cells and higher oncogenic potential.
Ultimately, the optimal vector is the one whose virological properties align precisely with the experimental or therapeutic objective-transforming viral biology from a constraint into a precision tool for genetic manipulation.
References
- Butt, Muhammad Hammad, et al. "Appraisal for the potential of viral and nonviral vectors in gene therapy: A review." Genes 13.8 (2022): 1370.
- Naldini, Luigi. "Gene therapy returns to centre stage." Nature 526.7573 (2015): 351-360.
- Wang, Dan, Phillip WL Tai, and Guangping Gao. "Adeno-associated virus vector as a platform for gene therapy delivery." Nature reviews Drug discovery 18.5 (2019): 358-378.
- Kuzmin DA, Shutova MV, Johnston NR, et al. The clinical landscape for AAV gene therapies. Nat Rev Drug Discov. 2021;20(3):173-174.

Your email address will not be published. Required fields are marked *